mRNA vaccination induces higher anti-RBD antibody levels than natural immunity against SARS-CoV-2
Many of the current serology tests for RBD have limitations on sensitivity, dynamic range, and unprecise or semi-quantitative3, 32, 33. To accurately determine the levels of anti-RBD antibody in SARS-CoV-2 infected individuals, we developed an electrochemiluminescence-based serology assay for exceptional sensitivity and dynamic range. The assay is based on two antibody-antigen interactions for maximum specificity (Fig. 1A). We purified recombinant wildtype (WT) RBD from transiently transfected 293 cells and labeled it with either biotin for antibody capture or ruthenium (Ru)-tag for antibody detection33. Two commercially available RBD monoclonal antibodies (Mab), Mab D001 and D003, were randomly selected (Sino Biological, Wayne, PA) and used as the calibrator (D003) and the reference standard (D001) for accurate quantification. Linear quantification range of at least 1,000-fold was achieved (Supplementary Fig. 1A). The assay was subsequently simplified from a three-step to a single incubation step without the loss of performance (Supplementary Fig. 1B), which would allow the rapid testing completed within one hour.
COVID-19 serology assay and antibody levels. (A) Diagram of the serology assay using electrochemiluminescence detection method. The capture RBD protein was attached to the assay plate via biotin and the detection RBD was conjugated to the voltage-sensitive ruthenium (Ru) molecule to emit light. (B) Quantitative RBD antibody data from sera samples of three cohorts: (1) samples on the first seropositive test, (2) convalescent donors with documented COVID-19, (3) donors without prior COVID-19 and completed two doses of mRNA vaccines. Negative control samples were collected from healthy donors prior to Jan. 2020. (C) The RBD antibody levels and time association in vaccinated donors (r = − 0.522, P = 0.0044). (D) Lack of RBD antibody levels and time association in COVID-19 convalescent donors (r = 0.234, P = 0.141).
The RBD serology assay was validated. In five consecutive tests, the average lower limit of quantification (LLoQ) was 0.57 ng/mL and the max LLoQ value of 0.97 ng/mL, with a linear quantification range of 1–1000 ng/mL (Supplementary Fig. 2A). Thus, 1 ng/mL was set as the LLoQ. When serum samples were adjusted for 1:10 dilution, the assay had a testing quantification range of 10–10,000 ng/mL (Supplementary Fig. 2A). In two sets of repeats of five experiments, the precision of quantification was 15.3 and 15.9%. The accuracy of the analysis was good with 100% spike recovery in two experiments, and intra-day CVs of 2.2 and 2.0%. Thus, the RBD serology assay has an LLoQ of 1 ng/mL and acceptable precision and accuracy.
The clinical validation study of the test was performed using 41 serum samples from 33 convalescent donors (Supplementary Table 1) with documented history of COVID-19 prior to September 1, 2020, months prior to the first reported alpha variant which contains RBD mutation N501Y and from 171 healthy donors collected before January 2020. None of the convalescent donors received COVID vaccines. The result revealed all negative samples were below the LLoQ (N = 171), whereas all convalescent donors were positive in the test (N = 41; Fig. 1B). The median RBD antibody level is 1.33 µg/mL with a large 170-fold range for RBD antibody levels in this small group of convalescent samples (range 27–4800 ng/mL; Fig. 1B). The RBD antibody assay has an area under curve (AUC) in Receiver Operator Characteristic (ROC) analysis of 1.00 (Supplementary Fig. 2B) with this groups of 41 samples from those with previously confirmed COVID-19 infections, thus, giving it an 100% sensitivity. It further has 100% specificity due to a lack of any quantifiable antibody levels in serum samples from pre-COVID-19 donors (N, 171).
In comparation with a commercially available spike S1 protein antibody test from Ortho Diagnostics, our RBD antibody test showed strong correlation with it (r = 0.679, P < 0.0001) (Supplementary Fig. 3). However, our assay exhibited a greater linear dynamic range at lower concentrations and improved sensitivity (41/41) over Ortho’s serology test (39/41) (Supplementary Fig. 3). Thus, our RBD antibody test is quantitative, and has exceptional sensitivity and a large dynamic range.
Clinical evaluation of RBD antibody levels in mRNA vaccinated and naturally infected individuals
We further evaluated the test in blood samples taken from those who went to the National Institutes of Health (NIH) Clinical Center for COVID-19 tests and were SARS-CoV-2 serology test positive for the first time. Similar levels of RBD antibody were detected in this newly diagnosed group (median, 0.74 µg/mL), which is not significantly different from those of convalescent individuals (P = 0.973) (N = 39, Fig. 1B). Interestingly, the blood from donors who completed two doses of mRNA vaccines (Pfizer or Moderna, N = 28) had much higher RBD antibody levels than that of the convalescent group and the newly diagnosed group (Fig. 1B, P < 0.0001). The median level of RBD antibody for the mRNA vaccine group was 22.3 µg/mL, which was 16.8-fold and 30.1-fold higher than that of the convalescent and newly diagnosed groups.
There were differences in age and in sampling time between the convalescent and immunized groups. The immunized group had a median age of 35.5 years which was different from that of the convalescent group at 59.0 (P < 0.0001) (Supplementary Table 1). However, the 17-fold differences in antibody levels between vaccinations and infections could not be explain by age alone, as there was an age overlap between the groups, and there was no age and antibody level association for both convalescent (P = 0.293) and immunized groups (P = 0.361) (Supplementary Fig. 4).
We examined the RBD antibody levels of the vaccine group and time association and noticed a correlation (r = − 0.522, P = 0.004; Fig. 1C). There was a difference in the antibody levels from samples taken within 2 months and at 6 months post second dose where the 6 months antibody levels were sharply lower (P = 0.001; Supplementary Fig. 5). In contrast, convalescent sera did not exhibit a correlative between time and antibody levels, with a median follow-up time of 207 days from the disease onset (r = 0.234, P = 0.141; Fig. 1D). The analysis of paired samples from same individuals in the convalescent group showed no change in antibody levels at two different time points (P = 0.396), whereas the paring was highly effective (r = 0.912, P = 0.0007; Supplementary Fig. 6). Hence, the data suggests that the antibody levels of convalescent sera did not decline significantly for 8 months post infections, whereas the ultrahigh RBD antibody levels achieved with mRNA vaccines could be subject to a more rapid decline.
Anti-RBD antibody concentration-dependent neutralization against RBD-ACE2 binding with COVID-19 antisera from natural immunity
To accurately quantify the ability of the antisera in neutralizing RBD and ACE2 binding, we developed an electrochemiluminescence based protein binding assay using recombinant RBD and ACE2 proteins as illustrated in Fig. 2A. When RBD was added to the assay wells, there was an excellent linear relationship between added free RBD and the luminescence signal from the RBD bound to ACE2 (r2 = 0.99; Fig. 2B). Therefore, the ACE2 binding assay provides a precise quantification of free RBD capable of binding to ACE2. The neutralization assay had an analytical precision of 6.8% (inter-day CV value, N = 3) using Mab D001 as the reference for neutralization assay. To clinically validate the sensitivity and specificity of the RBD-ACE2 binding assay, we analyzed the percentage of inhibition with the 41 COVID-19 convalescent sera and a comparable number of pre-COVID-19 control sera. The results showed that the 41 COVID-19 sera had a significantly higher inhibitory effect against RBD-ACE2 binding (P < 0.0001; Fig. 2C). The median levels of inhibition were 93% for the convalescent sera and 7% for the control sera. When comparing the convalescent sera with the negative controls, the antibody neutralization assay showed high sensitivity and specificity, with an AUC in ROC analysis of 0.986 (Supplementary Fig. 7), indicative of high sensitivity and specificity of the assay. The neutralization assay further has a clinical sensitivity of 90% and specificity of 97%, when a cutoff value (− 30%) was established based on the results of the pre-COVID-19 sera (Mean + 2X SD). Due to some non-specific inhibition of ACE2-RBD binding observed with pre-COVID-19 sera (Fig. 2C), additional work in better refining the cutoff and in determining the accurate sensitivity and specificity is needed. The neutralization assay further demonstrated similar consistency when compared with the RBD antibody test using paired convalescent serum samples taken at different time points, with a correlation coefficient for pairing r = 0.952 that was highly significant (P = 0.0001; Supplementary Fig. 8). Thus, not only the RBD-ACE2 receptor neutralization assay has excellent sensitivity and specificity for COVID-19 antisera, but also is precise with paired samples.
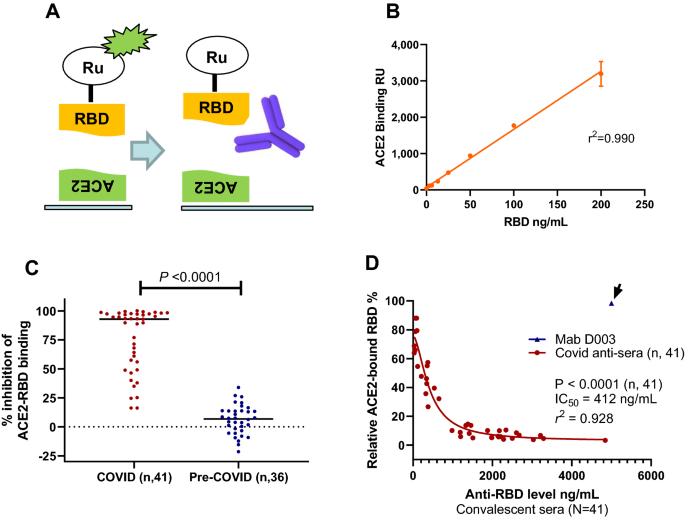
Neutralization assay, sensitivity, specificity, stability, and association with RBD antibody levels in COVID-19 convalescent sera. (A) Diagram of RBD and ACE2 biding assay and serum neutralization assay, where the neutralizing antibody prevents RBD from binding to ACE2. (B) Linearity of the RBD-ACE2 binding assay where increased RBD results in linear increase of binding signal (r2 = 0.990). (C) Neutralization data showing the convalescent sera have much higher neutralizing ability (P < 0.0001). (D) Serum neutralization assay results using COVID-19 convalescent sera showing strong correlation between anti-RBD levels and neutralizing activity (P < 0.0001, N = 41). The inhibitory concentration (IC50) shown was determined using nonlinear regression inhibitory model using RBD antibody levels in the sera, without accounting for a sixfold sample dilution.
From the eight donors with paired samples, it is also apparent that those with high anti-RBD levels (> 1000 ng/mL) showed stronger neutralization activity, whereas those with low anti-RBD levels (< 100 ng/mL) showed much lower neutralization activity (Supplementary Fig. 8). The complete analysis of COVID-19 convalescent sera for neutralization activity against RBD-ACE2 binding was performed. The results indicated there was an association between the anti-RBD antibody level and neutralization activity against RBD-ACE2 binding (correlative analysis, P < 0.0001; Fig. 2D). There was an extraordinary antibody dose-dependent neutralization activity using the nonlinear regression model with r2 = 0.928. The dilution adjusted (sixfold) and estimated 50% inhibitory concentration inhibition (IC50), based on the analyses of 41 samples, was 69 ng/mL. Five of the 41 convalescent sera had lower than 69 ng/mL RBD antibody, 4 of which were negative in the neutralization assay (< 30% inhibition, Fig. 2C). The neutralization assay was specific, as 5000 ng/mL of the calibrator monoclonal antibody D003, which bound to RBD well but exhibited no neutralization activity against RBD-ACE2 binding (Fig. 2D, data point shown with an arrow). As expected, RBD antibody concentration-dependent inhibition of RBD-ACE binding was confirmed with the second group of diagnostic serum samples (Supplementary Fig. 9). Thus, there is a strong RBD antibody concentration-dependent neutralization activity against RBD-ACE2 binding with two sets of COVID-19 antisera.
N501Y RBD significantly increased ACE2 binding and attenuated the neutralization ability of COVID-19 convalescent antisera
To investigate the ability of antisera from convalescent patient to recognize the B.1.1.7 N501Y variant (alpha), we purified N501Y RBD protein and labeled it with the Ru-tag for electro-chemiluminescence assay. We derived a scheme to compare the levels of antibodies against either the WT or N501Y RBD in the same samples (Supplementary Fig. 10). The results with the convalescent donor blood samples showed strong linear correlation between antibodies recognizing both the WT and the N501Y proteins (r2 = 0.927). Best-fit analysis revealed a slope of 1.09 (95% CI 0.99–1.19; Fig. 3A), suggesting the antisera from COVID-19 convalescent donors bind both the WT and N501Y RBD proteins equally well.
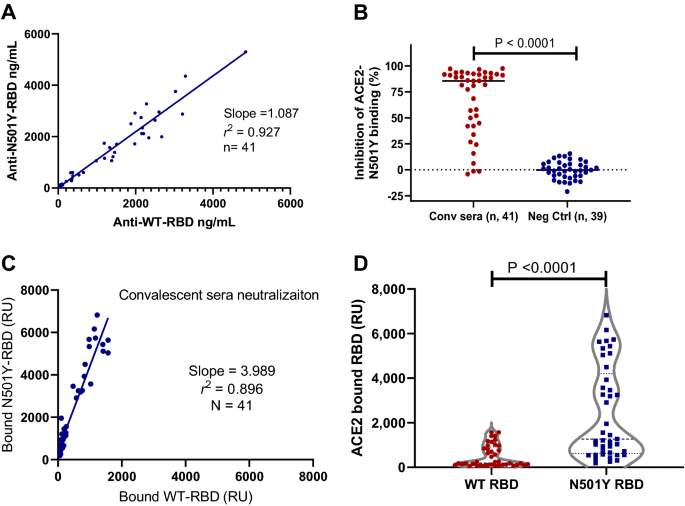
Comparison of antibody levels and neutralization activities against both the wildtype RBD and N501Y RBD proteins with convalescent sera. (A) Linearity of the anti-WT-RDB and anti-N501Y-RBD detected in the convalescent sera with a slope of 1.087 (r2 = 0.927). (B) Neutralization data from COVID-19 convalescent sera and negative controls. The convalescent sera had specific neutralizing activity against N501Y-RBD (P < 0.0001). (C) Linear regression analysis of the ACE2 bound the WT and N501Y RBD detected using the COVID-19 convalescent sera with a slope of 3.99 (r2 = 0.896, N = 41). (D) N501Y RBD have much higher absolute ACE2 binding than the WT RBD in the presence of neutralizing convalescent sera (P < 0.0001, N = 41).
We further analyzed the ability of COVID-19 convalescent antisera to neutralize the binding of the N501Y RBD to ACE2, as in the case of the WT RBD. The results showed the specific neutralization of the antisera from COVID-19 convalescent donors when compared with that of the pre-COVID-19 donor sera samples (P < 0.0001; Fig. 3B). The neutralization assay against RBD-N501Y and ACE2 binding was both very sensitive and specific with AUC of ROC analysis of 0.948 (Supplementary Fig. 11). There was further a strong linear correlation between neutralization activity against the WT and the N501Y RBD in ACE2 binding with a slope of 1.03 (r2 = 0.896, n = 41; Supplementary Fig. 12). Thus, COVID-19 antisera neutralize WT and N501Y RBD with an equal potency.
However, we observed dramatic differences in the ability of the WT and N501Y RBD to bind ACE2. Results from five consecutive experiments showed that N501Y RBD bound to ACE2 at an average of 5.1-fold higher rate than the WT RBD (range 4.1 to 6.1-fold). A representative result is shown (Supplementary Fig. 13). Thus, it can be concluded that N501Y RBD has a much higher affinity to ACE2. We further examined the absolute level of the WT and the N501Y RBD bound to ACE2 in the presence of COVID-19 antisera. While the antisera neutralized both the WT and N501Y RBD at a similar rate, there was four times (slope = 3.99, N = 41) more N501Y RBD bound to ACE2 in the presence of the convalescent antisera (Fig. 3C). This was further confirmed with the COVID-19 diagnostic sera (Supplementary Fig. 14). As more N501Y RBD has a higher affinity to ACE2, there were far more absolute amount of N501Y RBD bound to ACE2 than the WT in the presence of COVID-19 convalescent antisera (P < 0.0001, N = 41; Fig. 3D). Thus, natural immunity from the original SARS-CoV-2 infections could not consistently provide sufficient neutralization against N501Y RBD variant from binding to the cellular ACE2 receptor.
mRNA vaccination results in much more effective neutralization than natural immunity against N501Y RBD from binding to ACE2
To further determine the difference between natural immunity and mRNA vaccination, we selected five samples that had median levels of anti-RDB antibody of each group, and performed dilutions and neutralization studies against N501Y binding to ACE2. The results showed that dilution factors to IC50 were 25.8 and 402.0 for convalescent and mRNA vaccinated blood samples (Fig. 4A,B), a difference of 15.6-fold. This difference in neutralization is consistent with that the mRNA vaccinated blood had 16.8-fold higher anti-RBD antibody than the convalescent blood (Fig. 1B). Thus, the mRNA vaccinated blood is far more effective in neutralizing the high affinity N501Y RBD from binding to ACE2.
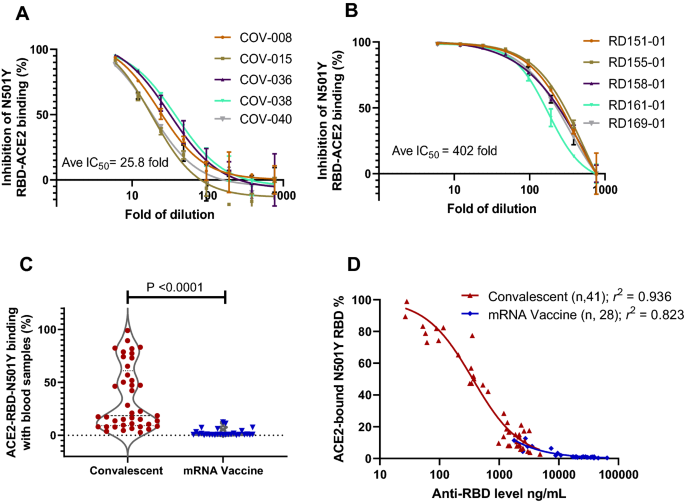
mRNA vaccine far more effective than natural immunity in neutralizing N501Y RBD mutant in ACE2 binding. (A) Dilution study to determine the IC50 of representative convalescent samples with median levels of anti-RBD antibodies. (B) Dilution study to determine the IC50 of representative vaccinated samples with median levels of anti-RBD antibodies. (C) Vaccinated blood samples more effective than convalescent ones to inhibit N501Y RBD in ACE2 binding (P < 0.0001). (D) Antibody concentration-dependent inhibition of N501Y RBD and ACE2 binding with blood samples from natural immunity (r2 = 0.926) and mRNA vaccination (r2 = 0.823).
When tested for neutralizing N501Y RBD against ACE2 binding, the mRNA vaccinated blood was far more effective compared to convalescent samples to achieve minimum ACE2 binding (P < 0.0001; Fig. 4C). There were 31.7% convalescent and 0% of vaccinated samples retained over 50% N501Y RBD and ACE2 binding, a difference that is highly significant. Detailed examination of antibody concentration dependent neutralization revealed strong correlations between RBD antibody levels and neutralization activities for both sample groups (r2 = 0.94 and r2 = 0.82; Fig. 4D). Thus, the substantially elevated antibody levels in the mRNA vaccine group appeared to be the primary driver of better neutralization activities against the high affinity N501Y RBD. Similarly, improved efficacy with the vaccinated blood was also observed against the WT RBD and ACE2 binding (Supplementary Fig. 15). Therefore, the data indicates the insufficiencies of natural immunity, and the superiority of mRNA vaccination in neutralizing RBD and ACE2 binding, particularly against SARS-CoV-2 variants with increased affinity to their cell receptor.